113568
论文已发表
注册即可获取德孚的最新动态
IF 收录期刊
- 3.4 Breast Cancer (Dove Med Press)
- 3.2 Clin Epidemiol
- 2.6 Cancer Manag Res
- 2.9 Infect Drug Resist
- 3.7 Clin Interv Aging
- 5.1 Drug Des Dev Ther
- 3.1 Int J Chronic Obstr
- 6.6 Int J Nanomed
- 2.6 Int J Women's Health
- 2.9 Neuropsych Dis Treat
- 2.8 OncoTargets Ther
- 2.0 Patient Prefer Adher
- 2.2 Ther Clin Risk Manag
- 2.5 J Pain Res
- 3.0 Diabet Metab Synd Ob
- 3.2 Psychol Res Behav Ma
- 3.4 Nat Sci Sleep
- 1.8 Pharmgenomics Pers Med
- 2.0 Risk Manag Healthc Policy
- 4.1 J Inflamm Res
- 2.0 Int J Gen Med
- 3.4 J Hepatocell Carcinoma
- 3.0 J Asthma Allergy
- 2.2 Clin Cosmet Investig Dermatol
- 2.4 J Multidiscip Healthc

在脓毒性休克儿童中通过使用生物信息学分析确定关键基因和途径
Authors Yang J, Zhang S, Zhang J, Dong J, Wu J, Zhang L, Guo P, Tang S, Zhao Z, Wang H, Zhao Y, Zhang W, Wu F
Received 16 November 2017
Accepted for publication 30 May 2018
Published 14 August 2018 Volume 2018:11 Pages 1163—1174
DOI https://doi.org/10.2147/IDR.S157269
Checked for plagiarism Yes
Review by Single-blind
Peer reviewers approved by Dr Justinn Cochran
Peer reviewer comments 3
Editor who approved publication: Dr Joachim Wink
Background and hypothesis: Sepsis is still one of the reasons for serious infectious diseases in pediatric intensive care unit patients despite the use of anti-infective therapy and organ support therapy. As it is well-known, the effect of single gene or pathway does not play a role in sepsis. We want to explore the interaction of two more genes or pathways in sepsis patients for future works. We hypothesize that the discovery from the available gene expression data of pediatric sepsis patients could know the process or improve the situation.
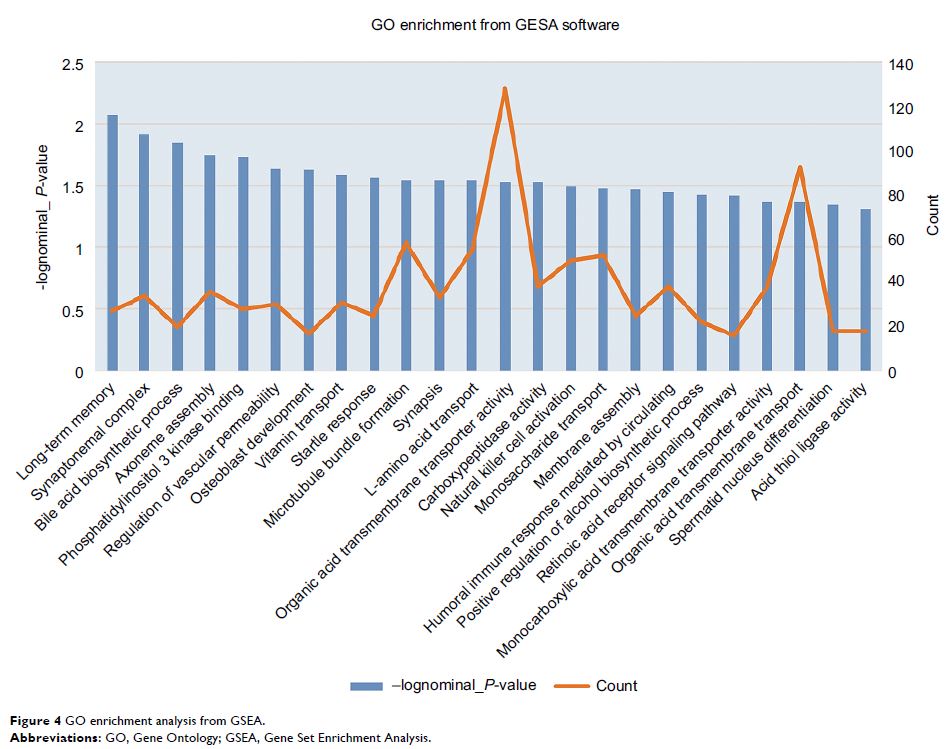
Methods and results: The gene expression profile dataset GSE26440 of 98 septic shock samples and 32 normal samples using whole blood-derived RNA samples were generated. A total of 1,108 upregulated and 142 downregulated differentially expressed genes (DEGs) were identified in septic shock children using R software packages. The Gene Ontology (GO) enrichment and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway were analyzed using DAVID software; Gene Set Enrichment Analysis method was also used for enrichment analysis of the DEGs. The protein–protein interaction (PPI) network and the top 10 hub genes construction of the DEGs were constructed via plug-in Molecular Complex Detection and cytoHubba of Cytoscape software. From the PPI network, the top 10 hub genes, which are all upregulated DEGs in the septic shock children, were identified as GAPDH , TNF , EGF , MAPK3 , IL-10 , TLR4 , MAPK14 , IL-1β , PIK3CB , and TLR2 . Some of them were involved in one or more significant inflammatory pathways, such as the enrichment of tumor necrosis factor (TNF) pathway in the activation of mitogen-activated protein kinase activity, toll-like receptor signaling pathway, nuclear factor-κB signaling pathway, PI3K-Akt signaling pathway, and TNF signaling pathway. These findings support future studies on pediatric septic shock.
Keywords: pediatric septic shock, microarray, differentially expressed gene, bioinformatics analysis